On an early June afternoon, Michael Linderman and I sat in our respective offices, our faces connected as they often are through a Zoom video call. Mine in Michigan, his in Vermont. I was eager to chat—I had only once before known someone to attend Harvey Mudd College, one of the seven contiguous Claremont Colleges huddled together in an otherwise sleepy Los Angeles suburb. I had heard stories from the other Mudd alumni I know about pranks pulled against their Caltech rivals (Google “Caltech Cannon Heist” if you want to learn more), and I wanted to know if Michael – SoundRocket customer, enigma – had been involved.
He wasn’t. Darn
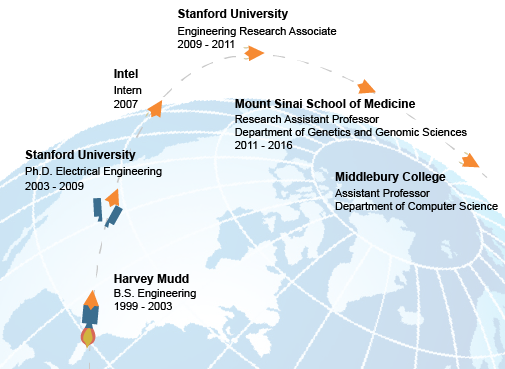
It’s no secret that this profile feature is named “Customer Trajectory” because it is, of course, about SoundRocket customers. After spending time getting to know Michael better, he was a clear choice to be our first contribution to this new feature of our blog, as the path that he has taken from his days at Harvey Mudd to his current Assistant Professor and Independent Consultant role has followed a trajectory that I don’t suppose many would have predicted.
I enjoyed getting to know Michael better, as I hope you do as well. . .
Q: So are you a scientist, and engineer, or something else?
A: I think the answer to your question is, a little bit of both. Increasingly, I find myself doing discovery-oriented work, but I think the reality is, a lot of that is influenced by the engineering background and is more engineering style work. The bioinformatics methods development is more akin to engineering than it might be to discovery-oriented science, where there is an ultimate discovery application, but in the near term, it’s focused on developing new methods and tools to enable that discovery.

Q: Who was the first scientist or engineer who had an impact on your life?
A: My lego set, as cliché as that sounds. I think I probably wrote the most clichéd college admissions essay. I can only imagine how tired they must be from reading about Lego’s. But it was pretty clear that I liked building stuff, and that’s what I wanted to do. I just knew that’s what I liked doing.
Q: Where did you grow up?
A: Washington, D.C. (Interviewers note: Wow, I missed out in a great follow-up here. “Did your father work for the CIA? Was your mother a Senator?” Oh well, we will never know.)
Q: Tell me something that we don’t know about you, maybe that would be a surprise to hear.
A: A surprise. I’m trying to think if there’s stuff that the students don’t know. [Long Pause] No, I don’t think I have a lot of mystery about me.
Q: Have you ever played hockey?
A: Nope. Don’t really understand the rules either, to be honest, other than the basics about goal scoring, maybe. But I enjoy the experience, and enjoy seeing the students that I have in class out on the ice.
Q: So you transitioned from computer science to working in a hospital. How did that happen?
A: Yea, well, I started spending more time with the immunology crew at Stanford, mostly to write software. The next two or three papers (after my first) were all in more engineering venues, but I kept working for them. I really liked the biology aspect and the applications to human health. It felt very important and significant. It also passed the Mom test in a way that working on compilers did not. When I went home for Thanksgiving and talked about what I did, I got glazed eyes when I spoke of compilers. But the life sciences stuff—immunology and neural-prosthetics—that produced a much more interested response.
I think I also realized it wasn’t really a viable long-term activity to just write software in an academic context. If I wanted to have a good experience, I needed to learn more about what the users were doing and the problems they were trying to solve so that I could become part of the science.
Q: Do you still write software?
A: Actually, just before you called, I was working on a method for trying to improve the accuracy of detecting a larger variance in sequencing data. I really like it, honestly. And I have students. This summer I have two students working with me on programming tasks. When I teach, more computer science oriented projects are needed to be amenable to the students.
Q: What big questions (in your field or not) really attracts you today?
A: Well, that would certainly be deep learning. That has applications everywhere for all things, including genomic stuff. We spend a lot of time doing variant interpretation, that’s a highly manual process right now that’s very judgement driven. Deep learning would seem to be a natural fit for this type of process. And the students are fascinated by it.
Q: If you were promised ten years of funding and offered a seat on a rocket to Mars, would you go?
A: I think there are aspects of going to Mars that I would do very well at. I’ve eaten the same lunch every day for years. So I’m not at all bothered by having the same freeze dried peas for years on end. I’d probably do well in confined spaces too. (Interviewers note: I unfortunately cut Michael off, as I truly got excited to hear about his lunch confession, before he ultimately said yes or no to the voyage, but I sense he was leaning towards reporting that he would be happy to remain here on earth.)
Q: The same lunch every day for years? Tell me about this lunch!
A: PB and Jam, string cheese, yogurt, and apple and a granola bar.
Michael’s Stats:
Current / Recent SoundRocket Collaborations:
- PeopleSeq Study: https://www.genomes2people.org/research/peopleseq/
- LinkedIn: https://www.linkedin.com/in/michael-linderman-45a80a2/
First Publication:
- Linderman, Michael D., Jamison D. Collins, Hong Wang, and Teresa H. Meng. “Merge: a programming model for heterogeneous multi-core systems.” In ACM SIGARCH Computer Architecture News, vol. 36, no. 1, pp. 287-296. ACM, 2008. https://dl.acm.org/citation.cfm?id=1346318
Most Recent Publication:
- Zoltick, Emilie S., Michael D. Linderman, Molly A. McGinniss, Erica Ramos, Madeleine P. Ball, George M. Church, Debra GB Leonard et al. “Predispositional genome sequencing in healthy adults: design, participant characteristics, and early outcomes of the PeopleSeq Consortium.” Genome Medicine, 11, no. 1 (2019): 10. https://genomemedicine.biomedcentral.com/articles/10.1186/s13073-019-0619-9
Stay tuned for more SoundRocket customer profiles, right here on the LaunchPad blog.

